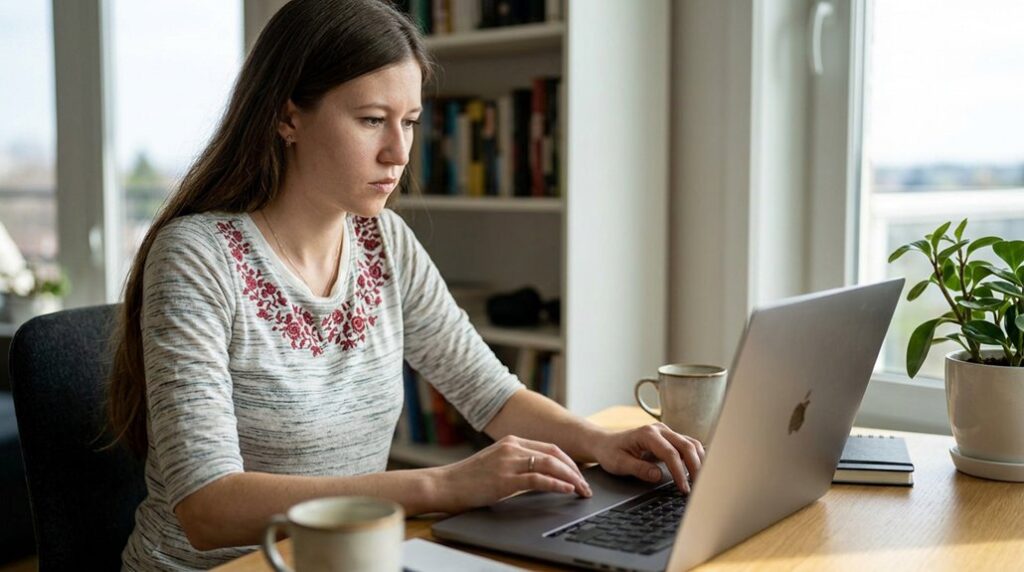
I recommend starting with R for Biologists, which walks readers through installing R and RStudio, then introduces variables, vectors, data frames, and ggplot2 with runnable code and clear checkpoints. Practical R for Biologists follows with real-world case studies and reproducible workflows. Biostatistics with R adds core statistical tests, while the Statistics for Biology and Health series offers a broader reference for readers who want to keep going.
Top R Programming Book Picks
R for Biologists: Learn R Programming from Scratch
Practical R for Biologists
Biostatistics with R
Statistics for Biology and Health
Coding for Biologists: Intro to Bioinformatics with Python
Computer Simulation and Data Analysis in Molecular Biology
Mixed Effects Models in Ecology with R
Statistics for Ecologists Using R and Excel
R Coding for Ecology
A Practical Guide to Ecological Modelling with R
More Details on Our Top Picks
R for Biologists: Learn R Programming from Scratch
Designed specifically for biologists who have never written a line of code, this book offers a gentle step-by-step pathway from installing R and RStudio to writing scripts that organize data in vectors, matrices, lists, and data frames. It explains core ideas in plain language and gives readers enough structure to work with CSV, Excel, or FASTA files without feeling rushed.
Practical R for Biologists
This is a strong next step for readers who want to use R on real datasets rather than only learn commands in isolation. The chapters are built around genuine scientific studies and guide readers through loading data, cleaning it, graphing it, and running statistical tests in a calm, practical way.
Biostatistics with R
If a reader needs to connect biological questions to formal statistical testing, this book is one of the better bridges. It pairs methods such as t-tests, ANOVA, regression, generalized linear models, and classification trees with R code and realistic case studies, which helps readers interpret results instead of only generating them.
Statistics for Biology and Health
This series is best seen as a broad searchable reference rather than a single beginner manual. It is useful for researchers who want deeper coverage of survival analysis, censored data, multivariate methods, and other specialised statistical topics while still working within an R-oriented biology context.
Coding for Biologists: Intro to Bioinformatics with Python
This is not an R book, but it makes a useful comparison point for readers who may eventually need Python for sequence analysis and bioinformatics. It starts with variables, loops, and functions, then moves into DNA, RNA, and protein sequence analysis in a way that remains approachable for complete beginners.
Computer Simulation and Data Analysis in Molecular Biology
This book is useful for readers who want to move from wet-lab intuition toward deterministic and stochastic modelling. It helps explain how computer simulations and statistical analysis can work together in molecular biology and biophysics, while keeping the discussion grounded enough for learners without strong computer-science backgrounds.
Mixed Effects Models in Ecology with R
For ecologists who need to analyse grouped, repeated, or hierarchical data, this is one of the most useful specialist books on the list. It combines theory with real ecological datasets and runnable R code, which makes it easier to see how mixed-effects models fit actual field data rather than abstract examples.
Statistics for Ecologists Using R and Excel
This is a practical bridge for readers who still manage much of their work in spreadsheets but want stronger statistical workflows. It is especially useful for ecologists who are moving from familiar Excel-based habits into more reproducible R-based analysis without abandoning the tools they already know.
R Coding for Ecology
This is better suited to readers who already know basic R and want to start applying it to ecological packages, datasets, and biodiversity workflows. It appears strongest as a next-step book rather than a first introduction, especially for researchers who want a broader overview of ecology-focused R tools.
A Practical Guide to Ecological Modelling with R
Readers who want to build ecological simulations rather than only run statistical tests will likely find this one especially useful. It combines theory with hands-on R code and covers models such as Lotka-Volterra, matrix models, lattice models, and decision models in a way that supports steady skill-building.
Factors to Consider When Choosing R Programming Books for Biologists
Audience Level and Prerequisites
The first thing to check is whether the book actually matches the reader’s current skill level. A title that assumes prior statistics, command-line confidence, or coding background can feel unnecessarily hard for a beginner. Books that clearly state their pace, prerequisites, and intended level are usually much easier to learn from.
Domain-Specific Examples
Examples built around biological data make the concepts stick much faster. It helps when a book shows how to handle CSV, Excel, FASTA, or ecological datasets, then walks through the entire workflow from import to cleaning, visualisation, and analysis using familiar biological questions.
Hands-On Code Exercises
Runnable code matters. The most useful books give short code-along tasks, project-style examples, and checkpoints so the reader can confirm that the output is correct. This makes troubleshooting easier and helps build confidence much faster than passive reading alone.
Visualization Tool Coverage
Good R books should cover not only statistical analysis but also visual communication. Practical guidance on ggplot2, base R graphics, and data reshaping with tidyverse tools is valuable because biology students and researchers often need to turn raw results into clear, publication-ready graphs.
Data Import Flexibility
Biological data rarely arrives in a single clean format. It is helpful when a book explains how to read spreadsheets, CSV files, sequence formats, databases, or web-based data sources, while also showing how to deal with missing values, mixed data types, and import errors without frustration.
Advanced Biological Applications
For more advanced readers, a stronger book may go beyond basic plotting and testing into genomics, transcriptomics, mixed-effects models, ecological simulations, or Bioconductor workflows. These topics are not essential for every beginner, but they matter if the goal is long-term scientific work rather than introductory exposure.
Final Thoughts
The best R book depends on what the reader needs right now. Some books are gentle entry points that teach the language from the ground up, while others are stronger for statistics, ecological modelling, or advanced biological applications. For most biology students and early-career researchers, one beginner-friendly foundation book plus one more specialised follow-up is often the most useful combination.
If you’re wondering how to turn your coursework and volunteer work into a real opportunity, read my step-by-step guide on how to get a paid biology internship or job with no experience.

Erzsebet Frey (Eli Frey) is an ecologist and online entrepreneur with a Master of Science in Ecology from the University of Belgrade. Originally from Serbia, she has lived in Sri Lanka since 2017. Eli has worked internationally in countries like Oman, Brazil, Germany, and Sri Lanka. In 2018, she expanded into SEO and blogging, completing courses from UC Davis and Edinburgh. Eli has founded multiple websites focused on biology, ecology, environmental science, sustainable and simple living, and outdoor activities. She enjoys creating nature and simple living videos on YouTube and participates in speleology, diving, and hiking.
🌿 Explore the Wild Side!
Discover eBooks, guides, templates and stylish wildlife-themed T-shirts, notebooks, scrunchies, bandanas, and tote bags. Perfect for nature lovers and wildlife enthusiasts!
Visit My Shop →